Abstract
Introduction
Bladder carcinoma (BC) initiates from the transitional epithelium of the bladder, and its invasion indicates poor prognosis and clinical outcome. 1 The highly invasive feature of BC specifically emphasizes the importance of comprehension of the molecular mechanisms underlying the control of BC metastasis.
Nuclear receptor coactivator 1 (NCOA1), also known as steroid receptor coactivator-1 (SRC-1), is a transcriptional coregulatory protein which is made up of several nuclear receptor–interacting domains and an intrinsic histone acetyltransferase activity.2–4 Intracellularly, NCOA1 is recruited to the specific sites for DNA promotion in a ligand-activated nuclear receptor–dependent manner. 4 NCOA1 has been found upregulated in a variety of cancers and are associated with tumor growth, invasion, and metastasis.5,6
MicroRNAs (miRNAs) are non-coding small RNAs that control some genes post-transcriptionally, through targeting the 3′-untranslated region (3′-UTR) of target messenger RNA (mRNA).7,8 Importantly, miRNAs have been found to control carcinogenesis and cancer progression.9–11 Specifically, the miRNAs have been found to play a critical role in the tumorigenesis of BC.12–19 However, no reports show that miRNAs may control NCOA1 during tumorigenesis. However, among all miRNAs, the function of miR-223-3p has been rarely studied and has been associated with the carcinogenesis of lung cancer, 20 prostate cancer, 21 and hepatocellular carcinoma. 22
Here, we examined the expression of NCOA1 and miR-223-3p in BC tissues and studied the association of miR-223-3p with prognosis of the patients. We further showed the regulatory relationship between miR-223-3p and NCOA1 using bioinformatics methods and proved it using luciferase-reporter assay. We then overexpressed miR-223-3p or inhibited miR-223-3p in two commonly used BC cell lines in vitro and studied their effects on NCOA1 activation and BC cell invasion.
Materials and methods
Experimental protocol approval
All experimental protocols were approved by the Research Bureau of First Affiliated Hospital of Nanchang University. All mouse experiments were approved by the Institutional Animal Care and Use Committee at First Affiliated Hospital of Nanchang University (Animal Welfare Assurance). The methods regarding animals and human specimens were carried out in “accordance” with the approved guidelines. Informed consent was obtained from all subjects.
Patient specimens
Surgical specimens from 24 BC patients (all Stage IV) and matched adjacent non-tumor bladder tissues (NT) were obtained postoperatively in First Affiliated Hospital of Nanchang University from 2011 to 2015. Informed consent was obtained from all subjects. All patients provided signed agreement for the resected tissue to be used for scientific research. The histology of the resected BC specimens and control tissue was confirmed independently by senior pathologists. All patients were followed up for 60 months.
BC cell culture and transfection
Human BC cell lines T24 and RT4 were both purchased from American Type Culture Collection (ATCC, Manassas, VA, USA) and have been widely used in BC research. T24 was generated from an 81-year-old female Caucasian, 23 and RT4 was generated from a 65-year-old male Caucasian. 24 Both cell lines were cultured in RPMI1640 medium (Invitrogen, Carlsbad, CA, USA) supplemented with 15% fetal bovine serum (FBS; Sigma-Aldrich, St Louis, MO, USA) in a humidified chamber with 5% CO2 at 37°C. MiRNAs mimics (miR-223-3p), miRNAs antisense oligonucleotides (as-miR-223-3p), null sequence, NCOA1, and short-hairpin small interfering RNA for NCOA1 (shNCOA1) were purchased from OriGene (Beijing, China). The transfection was performed with 50 nmol/L plasmids, using Lipofectamine 2000 (Invitrogen). The transfection efficiency was more than 95%, based on the expression of a green fluorescent protein (GFP) reporter.
Transwell cell invasion assay
Cells (104) were plated into the top side of polycarbonate transwell filter coated with Matrigel in the upper chamber of the BioCoat™ Invasion Chambers (Becton-Dickinson Biosciences, Bedford, MA, USA) and incubated at 37°C for 22 h. The cells inside the upper chamber with cotton swabs were then removed. Migratory and invasive cells on the lower membrane surface were fixed, stained with hematoxylin, and counted for 10 random 100× fields per well. Cell counts are expressed as the mean number of cells per field of view. Five independent experiments were performed, and the data are presented as mean ± standard deviation (SD).
MiRNA targets and 3′-UTR luciferase-reporter assay
MiRNAs targets were predicted with the algorithms TargetScan, as previously described. 25 The candidates were analyzed for context+ score, which is the sum of the contribution of six features (including site-type contribution, 3′ pairing contribution, local adenine/uracil (AU) contribution, position contribution, target-site abundance (TA) contribution, and seed-pairing stability (SPS) contribution). The NCOA1 3′-UTR reporter plasmid (pRL-NCOA1) and the NCOA1 3′-UTR reporter plasmid with a mutant at miR-223-3p binding site (pRL-NCOA1-mut) were purchased from Creative Biogene (Shirley, NY, USA). BC cells were collected 36 h after transfection for dual-luciferase-reporter assay (Promega, Fitchburg, WI, USA), according to the manufacturer’s instructions.
Quantitative reverse transcription polymerase chain reaction
Total RNA was extracted from resected tissue specimens or from the cultured BC cells, using miRNeasy Mini Kit (Qiagen, Hilden, Germany). Quantitative reverse transcription polymerase chain reaction (RT-qPCR) was performed in duplicates using QuantiTect SYBR Green PCR Kit (Qiagen), with the primers designed by Qiagen. A 2−ΔΔCt method was used to analyze and quantify the transcript levels. Values of gene transcripts were first normalized against housekeeping gene α-tubulin and then compared to the experimental controls.
Western blot
The protein was extracted from the cultured cells and homogenized in radioimmunoprecipitation assay (RIPA) lysis buffer (Sigma-Aldrich) on ice. The supernatants were collected after centrifugation at 12,000 ×
Statistical analysis
The SPSS 18.0 statistical software package was used to analyze data in this study. All values are depicted as mean ± SD and are considered significant if p < 0.05. A one-way analysis of variance (ANOVA) method with a Bonferroni correction, followed by Fisher’s exact test, was applied. Bivariate correlations were calculated by Spearman’s rank correlation coefficients. Kaplan–Meier analysis was used to analyze patients’ survival.
Results
Association of miR-223-3p levels in BC specimens with prognosis of the patients
The levels of NCOA1 and miR-223-3p in 24 pairs of resected BC tissues (Stage IV) and adjacent NT were measured by western blot and RT-qPCR, respectively. BC specimens contained significantly higher levels of NCOA1 (Figure 1(a)) and significantly lower levels of miR-223-3p (Figure 1(b)). We then performed a correlation test and detected a strong inverse correlation between NCOA1 and miR-223-3p (Figure 1(c), γ = −0.73, p < 0.0001, N = 24). In order to find out the clinical significance, these patients were followed up for 60 months. The median value for miR-223-3p in these patients was used as the cutoff point for separating miR-223-3p-high cases (n = 12) from miR-223-3p-low cases (n = 12). Kaplan–Meier curves were analyzed, showing that patients with low miR-223-3p in BC tissue had a significantly worse 5-year survival than those with high miR-223-3p in BC tissue (Figure 1(d)). These data suggest that low miR-223-3p levels in BC specimens may associate with poor prognosis.
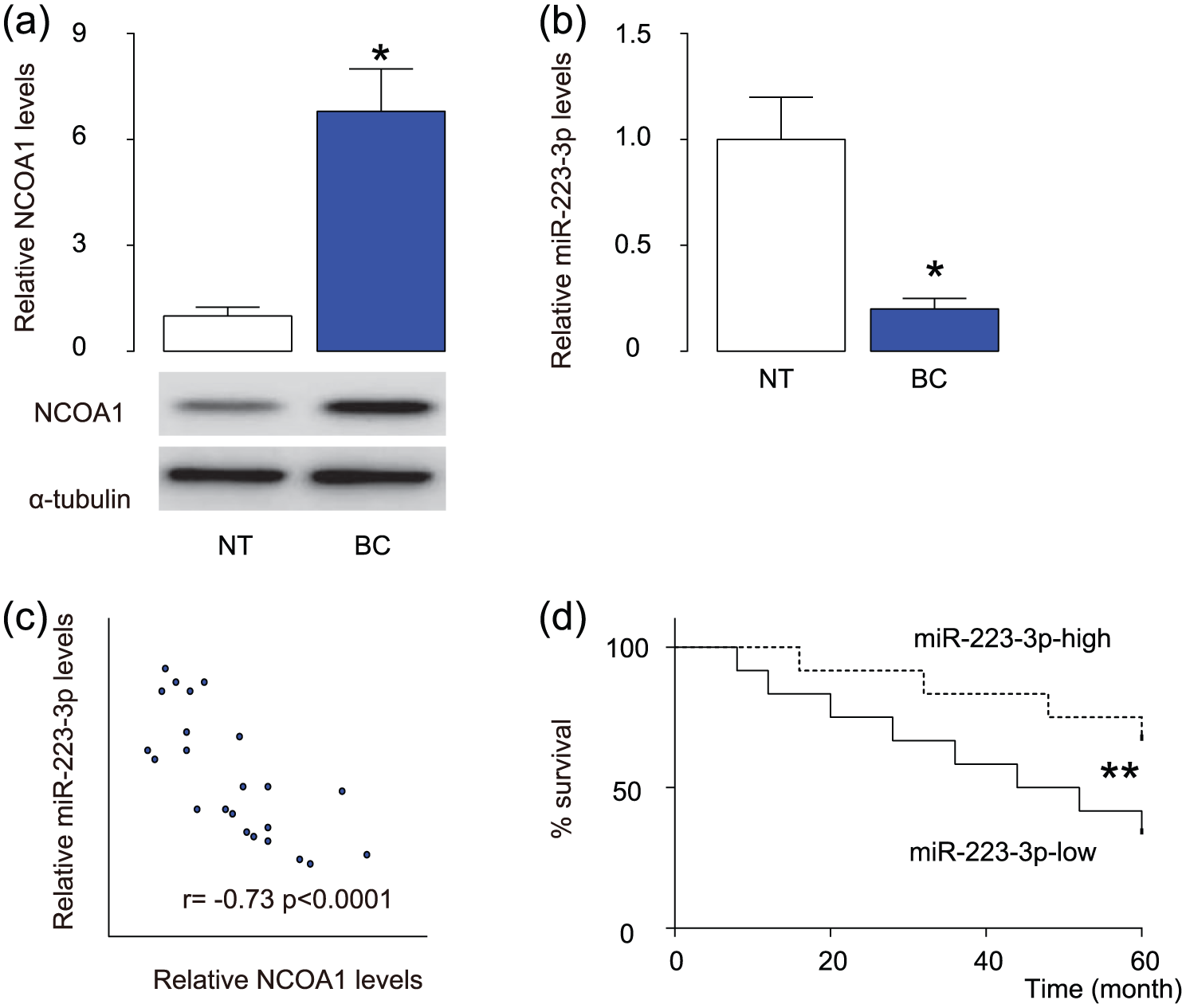
Low miR-223-3p levels in BC specimens associated with poor prognosis. (a)–(c) The levels of NCOA1 and miR-223-3p in 24 pairs of BC tissues and adjacent non-tumor bladder tissues (NT) were measured by (a) western blot and (b) RT-qPCR. (c) A correlation test was performed between NCOA1 and miR-223-3p using the 24 BC specimens. (d) The 24 BC patients were followed up for 60 months. The median value of all 24 cases was chosen as the cutoff point for separating miR-223-3p-high cases (n = 12) from miR-223-3p-low cases (n = 12). Kaplan–Meier curves were performed to compare 5-year survival between two groups (*p < 0.05; **p < 0.01; N = 24).
MiR-223-3p targets NCOA1 to inhibit its protein translation in BC cells
Next, we examined miR-223-3p and NCOA1 levels in different BC cell lines. Among these cell lines, we found that RT4 was a BC cell line expressing relatively high miR-223-3p and relatively low NCOA1, while T24 was a BC cell line expressing relatively low miR-223-3p and relatively high NCOA1 (Figure 2(a) and (b)). The functional binding of miR-223-3p to NCOA1 mRNA was predicted by bioinformatics algorithms (Figure 2(c)). Thus, we transfected T24 cells with miR-223-3p mimics (miR-223-3p; Figure 2(d)) and transfected RT4 cells with antisense for miR-223-3p (as-miR-223-3p; Figure 2(e)). The cells were also transfected with a null sequence as a control (null). The alteration of miR-223-3p levels in these cells was confirmed (Figure 2(d) and (e)). These miR-223-3p-modified BC cells were used to examine the functional binding of miR-223-3p to NCOA1 mRNA. The intact 3′-UTR of NCOA1 mRNA (NCOA1 3′-UTR), together with a 3′-UTR with mutant at miR-223-3p-binding site of NCOA1 mRNA (NCOA1 3′-UTR mut), was then cloned into luciferase-reporter plasmids. First, T24 cells were co-transfected with 1 µg miR-223-3p/null plasmids and 1 µg NCOA1 3′-UTR or NCOA1 3′-UTR mut plasmids (Figure 2(f)). Next, RT4 cells were co-transfected with 1 µg as-miR-223-3p/null plasmids and 1 µg NCOA1 3′-UTR or NCOA1 3′-UTR mut plasmids (Figure 2(g)). The results demonstrate that miR-223-3p specifically targets 3′-UTR of NCOA1 mRNA to inhibit its translation in BC cells.

MiR-223-3p targets NCOA1 to inhibit its protein translation in BC cells. (a) and (b) The levels of miR-223-3p (a) by RT-qPCR and NCOA1 and (b) by western blot in BC cell lines RT4 and T24, compared to BC tissue from patients. (c) Prediction of miR-223-3p-binding sites on NCOA1 mRNA by bioinformatics algorithms. (d) T24 cells were transfected with miR-223-3p mimics (miR-223-3p) or null as a control and examined for miR-223-3p levels. (e) RT4 cells were transfected with antisense for miR-223-3p (as-miR-223-3p) or null as a control and examined for miR-223-3p levels. (f) and (g) The intact 3′-UTR of NCOA1 mRNA (NCOA1 3′-UTR), together with a 3′-UTR with mutant at miR-223-3p-binding site of NCOA1 mRNA (NCOA1 3′-UTR mut), was then cloned into luciferase-reporter plasmids. (f) Luciferase activity was determined in T24 cells, which were co-transfected with 1 µg miR-223-3p/null plasmids and 1 µg NCOA1 3′-UTR or NCOA1 3′-UTR mut plasmids. (g) Luciferase activity was determined in RT4 cells, which were co-transfected with 1 µg as-miR-223-3p/null plasmids and 1 µg NCOA1 3′-UTR or NCOA1 3′-UTR mut plasmids (*p < 0.05; N = 5).
MiR-223-3p decreases NCOA1 protein but not mRNA in BC cells
The effects of miR-223-3p on NCOA1 were then analyzed in BC cells. Although the NCOA1 mRNA did not alter by miR-223-3p depletion in RT4 cells (Figure 3(a)), the NCOA1 protein was significantly increased by miR-223-3p depletion in RT4 cells (Figure 3(b)). Thus, miR-223-3p may decrease NCOA1 protein but not mRNA in BC cells.

MiR-223-3p decreases NCOA1 protein but not mRNA in BC cells. (a) and (b) The NCOA1 levels in miR-223-3p-depleted (and NCOA1-depleted) RT4 cells by (a) RT-qPCR and (b) western blot (*p < 0.05; N = 5).
Modification of miR-223-3p regulates BC cell invasion
The effects of miR-223-3p on the invasion of cultured BC cells were then investigated. We found that miR-223-3p depletion in RT4 cells significantly increased cell invasion in a transwell cell invasion assay (Figure 4(a) and (b)). Thus, suppression of miR-223-3p increases BC cell invasion.

Depletion of miR-223-3p abolishes RT4 cell invasion through NCOA1. (a) and (b) RT4 cell invasion by miR-223-3p depletion (and NCOA1 depletion) in a transwell cell invasion assay shown by (a) quantification and (b) representative images (*p < 0.05; N = 5).
MiR-223-3p regulates BC cell invasion through NCOA1
In order to figure out whether miR-223-3p may regulate BC cell invasion through NCOA1, we prepared plasmids for NCOA1 overexpression (NCOA1) and depletion (shNCOA1). First, RT4-as-miR-223-3p was further transfected with shNCOA1, resulting in decreases in NCOA1 mRNA (Figure 3(a)) and protein (Figure 3(b)) in these cells. Specifically, the effects of as-miR-223-3p on NCOA1 protein compromised the effects of shNCOA1 on NCOA1 protein, which explained the findings in BC cells transfected with both as-miR-223-3p and shNCOA1. We found that NCOA1 suppression abolished the effects of as-miR-223-3p expression on cell invasion in RT4 cells (Figure 4(a) and (b)). These data suggest that miR-223-3p may inhibit BC cell invasion through NCOA1 (Figure 5).

Schematic of the model. MiR-223-3p inhibits BC cell invasion through translational suppression of NCOA1.
Discussion
MiRNAs play demonstrative roles in the carcinogenesis in various cancers. In line with these notions, their participation in the BC progression has been widely reported.12–19 Previous studies have associated NCOA1 with tumor growth, invasion, and metastasis.5,6 In breast cancer, overexpression of NCOA1 was found to be associated with disease recurrence and resistance to endocrine therapy. Moreover, NCOA1 overexpression did not affect the morphology or tumor-forming capability of breast epithelial cells, but increased the number of circulating breast cancer cells and the efficiency of lung metastasis through colony-simulating factor 1 to enhance macrophage recruitment and metastasis. 6 In another report, 5 long-term NCOA1 knockdown on processes was relevant to metastasis formation. In vitro, Boyden chamber assays showed a strong decrease in migration and invasion upon NCOA1 knockdown, possibly through inhibition of protein kinase D1.
Inspired by these studies, here, we studied whether NCOA1 may be regulated by miR-223-3p in BC cells. Low levels of miR-223-3p in BC tissues were associated with poor survival rate in BC patients. Moreover, the levels of miR-223-3p and NCOA1 were inversely correlated. Then, we used a set of gain-of-function and loss-of-function experiments to show a regulatory relationship between miR-223-3p and NCOA1 in BC cells. In the promoter luciferase assay that showed that bindings of miR-223-3p to 3′-UTR of NCOA1 mRNA inhibited protein translation, there seemed to be a significant repression of the mutant 3′-UTR in the presence of miR-223-3p. Given that the mutant 3′-UTR should not bind miR-223-3p and yet did not have the same response as the wild-type 3′-UTR plus antisense, these data may suggest the presence of at least one cryptic miR-223-3p binding site. The large-scale changes in luciferase levels seemed unusual for miRNAs, which may be due to the importance of the binding site of miR-223-3p on the 3′-UTR of NCOA1 mRNA to its translation.
To summarize, our study here may provide evidence for using miR-223-3p as a novel target for treating BC and contribute to the understanding of molecular regulation of NCOA1-mediated BC cell invasiveness.
