Abstract
Impact Statement
Missing data are a common challenge in health analytics due to the technical and privacy challenges in collecting medical records. There is growing interest in imputing missing data in health datasets using deep learning. Existing deep learning–based imputation models have been commonly evaluated using root mean square error (RMSE), a predictive accuracy metric. In this article, we investigate the challenges of evaluating deep learning–based imputation models by conducting a comparative analysis between RMSE and evaluation metrics used in the statistical literature, including qualitative, predictive accuracy, statistical distance, and descriptive statistics metrics. To address these challenges, we design a new aggregated metric to evaluate deep learning–based imputation models called reconstruction loss (RL). We also present and evaluate a novel imputation evaluation methodology based on RL that researchers, system designers, and developers can use to develop predictive medical systems.
Introduction
Missing data are a common challenge when analyzing tabular datasets such as electronic medical records.1,2 One approach to handle missing data is imputation where the missing data are estimated using observed values in the dataset. There is growing interest in using deep learning to estimate the missing data. Deep learning imputation models include denoising autoencoders (DAEs 3 ), and generative adversarial nets (GANs 4 ). Deep learning models have three advantages over statistical imputation models such as logistic regression, decision trees, predictive mean matching (PMM), and sequential regression. 5 First, deep learning imputation can be used without making assumptions about the underlying distribution of the data. Next, missing data across multiple features can be estimated using a single imputation model. Finally, deep learning models can capture the latent structure of complex high-dimensional data (e.g. the correlation between demographics, medical history, and clinical outcomes in health records). 6
An important task in imputing missing data is evaluating the performance of the imputation models. 7 Imprecise models can produce misleading instances that impact the distribution of the groups being analyzed. The resulting discrepancies can impact a deep learning model’s performance. 8 The following motivating scenario demonstrates the importance of evaluating imputation models.
The authors of this article are currently developing a decision support system (DSS) using deep learning that assesses a patient’s risk from radiation exposure due to medical imaging (MI). 9 Our dataset has approximately 2.3 million imaging records from 340,525 patients over 10 years in four hospitals in Hamilton, Ontario, Canada. Due to technical and privacy challenges, we could access a subset of 18,875 DICOM (Digital Imaging and Communications in Medicine) headers. As a result, we need to impute the patients’ exposure in the remaining imaging records in the dataset using other features (e.g. body part). A common approach to estimating exposure is using mean values from the literature.10,11 However, a previous study 12 demonstrated that mean values from the literature under-estimate patients’ exposure. The resulting discrepancies had a cascading effect on the performance of the deep learning model.
A commonly used metric to evaluate the quality of a deep learning imputation model is the root mean square error (RMSE) which measures the difference between the imputed values and their corresponding actual values.3,4,13 RMSE is a performance evaluation metric. However, the goal of imputation in the statistical literature 14 is to ensure the imputed data meets the underlying properties of the dataset (e.g. data variability and distribution) rather than achieve the best prediction accuracy as in deep learning. 7
Preliminary results of this research are published in Boursalie
Materials and methods
In this section, we review existing imputation models, their evaluation methodology and metrics, and present our proposed evaluation methodology.
Imputation models
Consider a matrix
We can estimate missing data by drawing synthetic observations from the posterior distribution of the missing data, given the observed data and the process that generated the missing data. Formally, the posterior distribution is denoted as
Imputation models that estimate the posterior distribution of
Generative imputation models generate new instances from the posterior distribution of
Missing data can be estimated using single or multiple imputation.
20
Multiple imputation captures the uncertainty of the imputation model by performing
Evaluation methodology and metrics
The evaluation methodology proposed in the literature to assess the quality of an imputation model is as follows 21 :
In the statistical literature, the evaluation metrics (Step 4) can be qualitative (e.g. histogram, box, and density plots) or quantitative (e.g. predictive accuracy, statistical distance, and descriptive statistics) as shown in Table 1. Predictive accuracy metrics measure the difference between the imputed values and their corresponding actual values. RMSE is a predictive accuracy metric and is defined in equation (1) where
Summary of evaluation metrics.

RMSE: root mean square error; CDT: Cohen’s Distance Test; KL: Kullback–Leibler; IQR: interquartile range.
Unlike predictive accuracy metrics, statistical distance metrics such as Cohen’s Distance Test (CDT
22
) and

where
Descriptive statistics describe the characteristics of the distribution such as frequency, central tendency (mean, median, and mode), measures of variability (standard deviation and skewness), and position (quantile ranks). Normal distributions are described using mean and standard deviation. Non-normal distributions are described using median

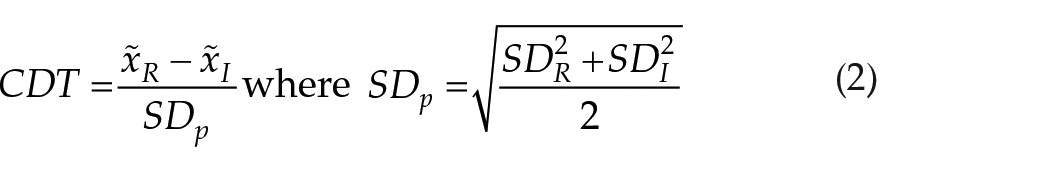

Existing proposals (MIDAS,
13
GAIN,
4
and VAE
3
) assessed the deep learning imputation models using the methodology proposed in Marshall
Proposed evaluation methodology
To address the limitations in the existing imputation evaluation methodology, 21 our proposed methodology needs to attend to the following requirements:
To address R1, we evaluate the imputation model’s performance on three metrics: (1) median, (2) skewness, and (3) IQR reconstruction. Median, skewness, and IQR were selected for this study because we are interested in imputing non-normal distributions. To address R2, we want to aggregate the median, skewness, and IQR performance. However, we cannot sum median, skewness, and IQR because each metric has different ranges and definitions. For example, a negative skew value represents a left-skewed distribution while a negative median represents a number in the dataset. To summarize the performance of multiple metrics, we need to do the following: (1) make all metric values positive, (2) compare the differences between the metric values for the imputed and target distributions instead of the values themselves, and (3) normalize each metric. Our proposed metric, reconstruction loss (

where

We are using weights
Figure 1 extends the existing imputation evaluation methodology with our proposed RL metric. First, we impute the missing data introduced into the dataset using the candidate imputation models. Second, we calculate the dataset properties (mean, skewness, and IQR) for the target and imputed distributions and perform data shifting to remove negative values, if necessary. Third, we calculate and plot the average

Proposed imputation evaluation methodology.
Results
In this section, we present the comparative analysis of qualitative, predictive accuracy, statistical distance, descriptive statistics, and our proposed RL metric to assess two deep learning imputation models (MIDAS and GAIN) and a regression-based imputation model (PMM) on two tabular datasets. MIDAS and GAIN represent non-generative and generative deep learning imputation models, respectively. We selected PMM as our benchmark model from the statistical literature when evaluating the deep learning models’ performance.
Data collection and processing
We evaluated the imputation models (PMM, MIDAS, and GAIN) on two tabular datasets: (1) MI and (2) Credit.
30
The MI dataset was collected in a retrospective study we performed of all medical scans from 1200 patients who received at least one low-dose MI scan (e.g. CT and XR) from four hospitals in Canada between May 2006 and May 2017. The patients were a stratified random sample representative of the target population in terms of gender, age of first scan, and body part scanned. The patients also had above-average cumulative ED
31
exposure. ED is a metric to estimate the uniform whole-body dose that has the same nominal radiation risk compared to the non-uniform exposure from MI.
31
Table 2 describes the characteristics of the MI dataset. Each patient’s medical history contains demographic, health, and imaging records. Demographic data include the patient’s age, sex, year, and month of the medical visit. Health records include diagnostic codes in the International Statistical Classification of Diseases and Related Health Problems (ICD-10-CA) format. Imaging records consist of modality (CT or XR) and body part scan (e.g. head) in the DICOM format. The ED exposure from MI is estimated using the methodology from Boursalie
Medical imaging and Credit dataset characteristics.

C: continuous and D: discrete features.
The Credit dataset 30 contains 30,000 banking records from clients at a Taiwanese bank between April and September 2005. Table 2 describes the characteristics of 29,206 clients who received or paid a bill between April and September 2005. Each client’s banking information includes the client’s sex, credit limit, education level, marital status, monthly bills, payments, and repayment status (bill paid on time or the amount of months payment was late). All continuous features are normalized using min–max normalization. Additional information on the Credit dataset is available at Yeh and Lien. 30
The MI and Credit datasets were selected for this study because they have continuous and discrete features with no missing data. The Credit dataset was also used to evaluate GAIN.
4
Unlike previous studies,4,13 to be consistent with the evaluation methodology,
21
we selected one target feature for each dataset to impute. We imputed ED
Evaluation procedure
We assessed the performance of PMM, MIDAS, and GAIN to impute missing data in the MI and Credit datasets. We introduced increasing proportions of data MCAR (2%, 4%, 8%, 10%, 20%, 40%, and 80%) in the target features (Table 2) after preprocessing. We validated (Supplemental Appendix A) that our missing data mechanism (MCAR) did not change the statistical properties of the train and test sets, so no data leakage (bias) occurred by preprocessing our dataset before introducing missing data. We selected the data proportions of MCAR to be consistent with previous studies.4,13,21 We imputed the missing data at each proportion using PMM, MIDAS, and GAIN. We took the mean results
All experiments were conducted on a 64-bit Windows 7 laptop with a 2.8 GHz Intel Xeon CPU and 16 GB RAM. The default PMM
5
architecture (50 epochs) was implemented using the open-source code. For MIDAS
13
and GAIN,
4
the default architectures (TensorFlow), loss functions (RMSE and cross-entropy), epochs (MIDAS: 20, GAIN: 10,000), optimizers (MIDAS and GAIN: Adam Optimizer), and learning rates (MIDAS: 1, GAIN: 1.5) were implemented using their open-source codes. Interested readers are referred to Lall and Robinson
13
and Yoon
Comparative analysis
Figure 2 shows a subset of the qualitative histogram results.

Histogram of
The imputation models’ RMSE performances imputing

RMSE (a and b), CDT (c and d), and JSDist (e and f) evaluation results for
Figure 3(c) shows the CDT performances for the
The imputation models’ JSDist performances imputing
The median metric results of the

Median (a and b), skewness (c and d), and IQR (e and f) descriptive statistics results for
The skewness metric results of the
The IQR metric results of the
Figure 5 shows a subset of the average RL performance for the

RL values for QT MIDAS (a), No-QT MIDAS (b), QT PMM (c), No-QT PMM (d), QT GAIN (e), and No-QT GAIN (f) imputation models for 80% missing

RL values for QT MIDAS (a), No-QT MIDAS (b), QT PMM (c), No-QT PMM (d), QT GAIN (e), and No-QT GAIN (f) imputation models for 80% missing
Discussion
Table 3 ranks each imputation model’s performance based on the qualitative, predictive accuracy, and statistical distance metrics for each dataset we investigated. The qualitative results were ranked based on a visual inspection of the mean and distribution reconstruction (Figure 2) across all runs. Based on the histogram (benchmark) results, the QT-PMM would be selected for imputation in both datasets across all missing data rates. However, the predictive accuracy metrics did not agree with the qualitative and statistical distance results. The No-QT-MIDAS would be selected for the MI dataset and the No-QT-PMM, No-QT-MIDAS, or QT-MIDAS models would be selected for the Credit dataset based on RMSE. Using CDT, the No-QT-PMM, QT-PMM, or No-QT-MIDAS model would be selected for the MI dataset and the No-QT-PMM, QT-PMM, No-QT-MIDAS, or QT-MIDAS would be selected for the Credit dataset. The JSDist ranking agreed with the histogram results. The qualitative results (Figure 2) provide an initial check of the imputation model’s performance
7
and provide context to the quantitative metrics. For example, the qualitative results demonstrate that the poor performance of the GAIN models is due to mode collapse. The qualitative results can also be reviewed by an expert to evaluate imputation models with similar performance. For example, a medical expert could review the qualitative results of the No-QT-PMM and QT-PMM models (Figure 3(e)). Our results demonstrate that the predictive accuracy metrics evaluate the imputation model’s ability to capture the mean of the target distributions. For example, RMSE directly compares the imputed estimates with the actual values rather than comparing the distributions (equation (1)). Similarly, CDT compares the means and standard deviations of the imputed and actual distributions (equation (2)). Interestingly, the imputation models that generate more normal distributions (MIDAS and no-QT models) minimized their RMSE and CDT (Figure 3(a) to (d)) without capturing the distribution of the target features. In fact, the PMM models that attempted to capture the distribution of the target features (Figure 2) had poorer RMSE and CDT performance (Figure 3(a) to (d)). In addition, the GAIN models demonstrate how competitive RMSE and CDT results (Figure 3(a) to (d)) can be achieved by imputing a single value due to mode collapse. Our results demonstrate how imputation models can achieve good predictive accuracy performance (Figure 3) without capturing the feature’s distribution (Figure 2). The
Imputation models’ ranked performances (1 best, 4 worst) based on the evaluation metrics.

ED: effective dose; PMM: predictive mean matching; MIDAS: Multiple Imputation with Denoising Autoencoders; GAIN: Generative Adversarial Imputation Nets; QT: quantile transform; RMSE: root mean square error; CDT: Cohen’s Distance Test; IQR: interquartile range; RL: reconstruction loss.
The descriptive statistics assessed different characteristics of the imputation model’s performance (median, skewness, and IQR). Interestingly, the deep learning–based model (QT MIDAS) competitive reconstruction performance was not captured when evaluating the models using the RMSE, CDT, and JSDist metrics. Our results demonstrate that previous studies that have evaluated deep learning imputation using predictive accuracy metrics4,13 may not capture the overall performance of their models. However, our findings also suggest that the performance metric should be selected based on the dataset size, distribution of features, and proportion of missing data. While descriptive statistic metrics provided us insights into the imputation model’s behavior on the ED dataset, the metrics were not as sensitive in evaluating imputation performance on the Credit dataset (Figure 4). Overall, our study demonstrated that qualitative, predictive accuracy, statistical distance, and descriptive statistics investigate different properties of reconstruction performance, and there is a need to aggregate performance between these metrics.
Our proposed evaluation methodology (Figure 1) successfully ranked the performance of the imputation models. Unlike the existing methodology (section “Materials and methods”), our methodology considered multiple aspects of the imputation model’s reconstruction performance (mean, skewness, and IQR). In addition, our methodology provides a mechanism for users to study the trade-offs between the reconstruction criteria. For example, the best balance between median, skewness, and IQR reconstruction was achieved by the MIDAS model. Unlike RMSE, CDT, and JSDist, our methodology did not penalize the MIDAS and PMM models for attempting to capture the distribution of the target features at the expense of mean and skewness reconstruction. Our methodology also provides a mechanism to incorporate expert opinion when selecting an imputation model beyond qualitative analysis.
In the statistical imputation literature, researchers impute datasets using multiple methods and investigate their impact on the downstream predictive model. Our proposed evaluation methodology provides researchers with a quantitative method to select the imputation models for further study. For example, we can investigate how the downstream deep learning model’s performance changes when imputing data using GAIN (mean imputation), PMM (IQR imputation), and MIDAS (median, skewness, and IQR imputation) models.
RL can be used to evaluate the imputation model’s performance on normal and non-normal target distributions. However, the dataset properties may determine the suitability of the evaluation metrics to assess the imputation models. For example, the statistical distance metric (JSDist) may better capture the difference between the target and imputed distributions for non-normal distributions compared to predictive accuracy metrics. Similar to statistical tests, 5 the choice of metric may depend on the dataset size. In our study, the metrics had similar performances (Figures 3 to 6) between datasets with different sizes (Table 2).
Our study exhibits some limitations. First, we investigated the imputation model’s performance on specific features (ED and age). In addition, data MAR and MNAR were not investigated. Finally, we investigated the default model architectures. Improved model architecture and training could impact performance evaluation.
Related work
An important component in deep learning is evaluating performance. Previous studies have evaluated various properties of the deep learning models’ performances such as accuracy, computational,
35
robustness,
36
privacy,
37
ethical,
38
and trust.
39
Increasingly, deep learning models are evaluated using multiple performance metrics.
35
Furthermore, benchmarks
40
are being developed to evaluate new models with prior work using open-source datasets (e.g. ImageNet
41
). There is also a growing body of literature surveying the strengths and limitations of evaluation metrics for deep learning19,42,43 to assist researchers and developers to select their evaluation metrics. For example, Borji19,44 and Thompson
In statistics, there is a large body of surveys on metrics that can be used to evaluate imputation models. For example, Deza and Deza
45
encyclopedia of distances provides an overview of available distance metrics for comparing distributions. Previous studies7,19 also provide guidelines for researchers to select their metrics to evaluate statistical imputation models. Nguyen
Despite advances in the deep learning and statistical literature, existing deep learning imputation models (MIDAS,
13
GAIN,
4
and VAE
3
) have been assessed using RMSE, a predictive accuracy metric. The studies showed the deep learning imputation models had competitive RMSE performance compared to statistical imputation models. However, deep learning imputation models can impute missing data in multiple features at once. As a result, Lall and Robinson
13
and Yoon
Conclusions
The existing evaluation methodology commonly used to assess deep learning–based imputation models lacks a mechanism to evaluate and investigate multiple aspects of the model’s reconstruction performance. To address this challenge, we proposed an evaluation methodology and an RL metric to assess deep learning–based imputation models. Our methodology ranks imputation models by their performance across multiple reconstruction properties such as median, skewness, and IQR. Our methodology also provides researchers with a mechanism to evaluate the trade-offs between reconstruction properties. We used our evaluation methodology to assess two deep learning imputation models on two tabular datasets. We also described the strengths and challenges of using evaluation metrics to assess deep learning–based imputation models.
Given these results, we are extending our proposed imputation evaluation methodology to rank the deep learning model’s imputation performance across multiple features with missing data. In addition, we will investigate how deep learning imputations perform for multiple features compared to statistical imputation models. Finally, we are investigating methods to improve visualizing the trade-offs between imputation performance metrics.
Supplemental Material
sj-docx-1-ebm-10.1177_15353702221121602 – Supplemental material for Evaluation methodology for deep learning imputation models
Supplemental material, sj-docx-1-ebm-10.1177_15353702221121602 for Evaluation methodology for deep learning imputation models by Omar Boursalie, Reza Samavi and Thomas E. Doyle in Experimental Biology and Medicine
Footnotes
Authors’ Contributions
Declaration of Conflicting Interests
Funding
Supplemental Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
