Abstract
Introduction
The interaction of biological entities such as genes, proteins and metabolites is of great interest in life science research and is increasingly important for systems biological approaches (Oltvai and Barabási (2002); Kitano (2002)). The interplay of different interactions is often represented by biological networks such as gene regulatory, protein interaction and metabolic networks. To investigate these complex and large networks different network analysis methods have been developed or employed from other fields of sciences (Junker and Schreiber (2008)). Centrality analysis, the ranking of network elements used to identify interesting elements of a network is one of these methods (Koschützki et al. (2005)). It is particularly useful to identify key players in biological processes. For example, it has been shown that highly connected vertices in protein interaction networks are often functionally important and the deletion of such vertices is related to lethality (Jeong et al. (2001)). Wuchty and Stadler applied three different types of centralities to metabolic, protein interaction and domain sequence networks (Wuchty and Stadler (2003)). Fell and Wagner discuss the possibility that metabolites with highest degree (i.e. highest number of connections) may belong to the oldest part of the metabolism (Fell and Wagner (2000)). However, it has also been shown that the degree of a vertex alone, as a specific centrality measure, is not sufficient to distinguish lethal proteins clearly from viable ones (Wuchty (2002)), that in protein networks there is no relation between network connectivity and robustness against amino-acid substitutions (Hahn et al. (2004)), and that for biological network analysis several centrality measures have to be considered (Wuchty and Stadler (2003); Koschützki and Schreiber (2004)).
To assist scientists in the exploration of biological networks, we discuss and compare different centrality measures. Some of them are already known in biological sciences, others are transferred from different fields of sciences such as social network analysis. We also show that it is useful to consider biological knowledge in network analysis and discuss motif-based centralities which have been specifically developed for gene regulatory networks.
Graphs and Centralities
A network is an informal description for a set of elements with connections between them. In a formal way a network is modelled as a mathematical object called graph. A directed graph
The degree
A subgraph of the graph
Small recurring subgraphs within a given graph are called motifs (Milo et al. (2002)). A motif
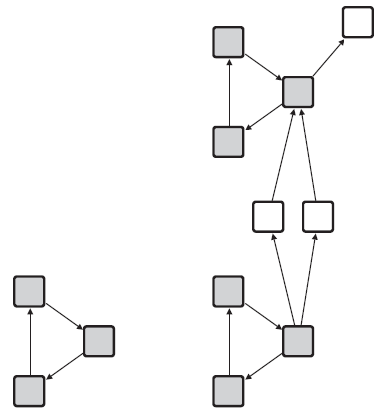
A motif and two matches of the motif in a graph.
Centralities in networks
Formally a centrality is a function
In the following sections we explain different centrality measures and show an example graph and the corresponding centrality values. We restrict our analysis to centrality measures which have been used to analyze biological networks or are used in our study in the second part of this paper. A comprehensive overview of different centrality measures was published in (Koschützki et al. (2005)).
Degree centrality
An obvious order of the vertices of a graph can be established by sorting them according to their degree. The corresponding centrality measure degree-centrality is defined as
Closeness centrality
Closeness-centrality uses information about the length of the shortest paths within a network; it uses the sum of the minimal distances of a vertex to all other vertices. The closeness-centrality is defined as the reciprocal of this sum:
Radiality and integration
Similar to the closeness measure are the centralities radiality and integration introduced by Valente and Foreman (1998). The computation of both centralities is based on the reverse distance matrix which is defined on the basis of the distance matrix
A vertex with a high radiality value can easily reach other vertices. A vertex with a high integration value is easily reachable from other vertices. Similarly to closeness both radiality and integration are shortest path based measures. In contrast to closeness which can be only computed for strongly connected networks, radiality and integration can also be computed for weakly connected or even unconnected networks.
Shortest path betweenness centrality
Shortest path betweenness centrality quantifies the ability of a vertex to monitor communication between other vertices. Every vertex that is part of a shortest path between two other vertices can monitor communication or flow between them. Counting how many such communications a vertex may monitor leads to an intuitive definition of a centrality: a vertex is central if it can monitor many communications between other vertices. In the following let
There are several studies investigating shortest path betweenness in biological networks. For an
Katz status index and PageRank
For the analysis of gene regulatory networks discussed in the second part two further centralities can be applied: the status index defined by Katz (1953) and the PageRank centrality (Page et al. (1998)) which is the algorithmic method behind the search engine Google. Both centralities are best described as computations performed on the adjacency matrix accompanied to the graph of interest. As we focus on the result of different centralities and their comparison we skip a lengthy formal definition here and refer to the literature for details (Katz (1953); Page et al. (1998); Koschützki et al. (2005); Koschützki (2008)).
Motif-based centralities
Given a graph
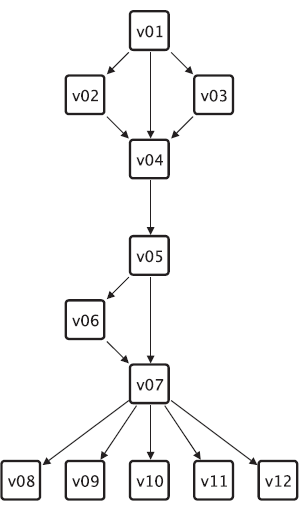
An example graph used to explain different centrality measures.
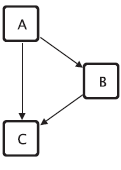
The FFL motif with roles.
Vertices of motifs may represent different functions. For example, in the gene regulatory network context three different functions of the vertices of the feed forward loop (FFL) motif as shown in Figure 3 can be identified: (1) the vertex at the top is the master regulator, this vertex regulates the other two vertices; (2) the vertex on the right side is the intermediate regulator, it is regulated by the master regulator and itself regulates together with the master regulator the vertex at the bottom; and (3) the vertex at the bottom of the drawing is regulated by both other vertices and is therefore called the regulated vertex. Such different functions of vertices within motifs are called roles and three roles can be assigned to the vertices of the FFL motif. The motif-based centrality with roles
Using the previously introduced concepts we can extend the motif-based centrality method further. By assigning the same role to similar vertices of a group of similar motifs we can establish a centrality based on a class (or group) of motifs. Consider, for example, a group of chains (see Fig. 4), where all vertices at the start of such chains have a similar characteristic (no incoming edges) and all vertices at the end have another similar characteristic (no outgoing edges). For gene regulatory networks several motif classes are known. For example, the regulatory chain motif class, as in the example above, consists of a set of chains of three or more regulators in which one regulator regulates another regulator, which in turn regulates a third one and so forth (Lee et al. (2002)). In the motif class single input motif (SIM) a set of vertices is exclusively regulated by a single vertex (Shen-Orr et al. (2002)). The motif-based centrality with classes
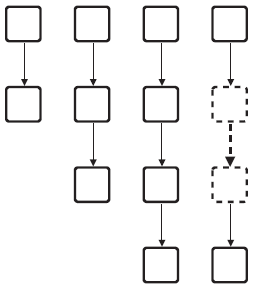
Several motifs of the chain motif class.
Several motifs have been studied in all kinds of biological networks. The best studied motif is the FFL motif which functional properties have been analyzed in detail theoretically and experimentally especially in gene regulatory networks (Mangan and Alon (2003); Mangan et al. (2003); Shen-Orr et al. (2002); Wall et al. (2005)). However, in these approaches only the occurrence of motifs is considered but motifs are not used to rank the genes.
Different motifs occurring in a human cellular signalling network were analysed by Awan et al. (2007). They discovered that genes which are related to cancer are enriched in the target vertices of several motifs and that cell mobility genes are enriched in the source vertices of motifs. For a gene regulatory network of
Example graph and centralities
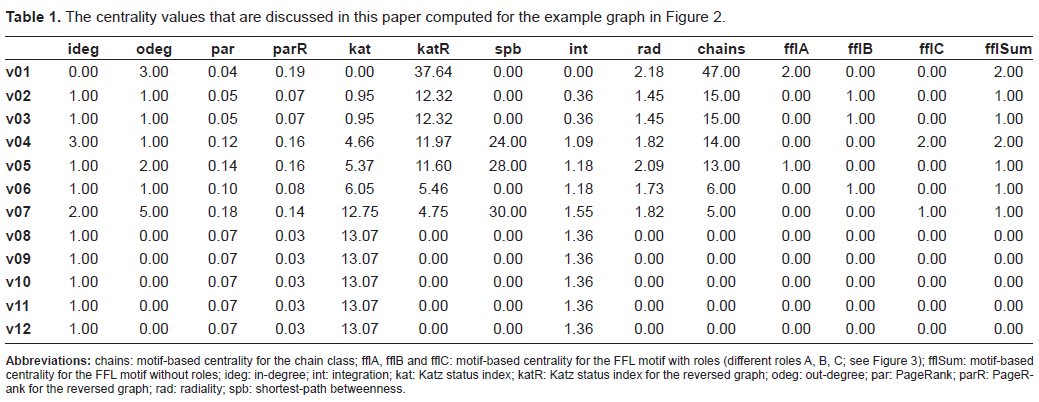
Figure 2 shows a small example graph and the corresponding Table 1 shows the centrality values for the centralities that are applicable to this graph.
The centrality values that are discussed in this paper computed for the example graph in Figure 2
Analysing Gene Regulatory Networks with Centralities
The applicability of specific centrality measures for the investigation of biological networks depends on the type of the particular network, and depending on the type of the network different centrality measures are used. Here we focus our analysis on gene regulatory networks.
As an example, we analyze centralities within the gene regulatory network (GRN) of
Comparison of different centralities for GRN
In this section, we compare different centrality measures that can be applied to GRNs. As GRNs are directed graphs that are not necessarily strong connected only the centralities degree, shortest-path betweenness, integration, radiality, Katz status index, PageRank and the different motif-based centralities can be applied. The centralities PageRank and Katz status index are sensible to the directionality of the edges and therefore we consider two variants of the graph, the original graph and the graph with all edge directions reversed.
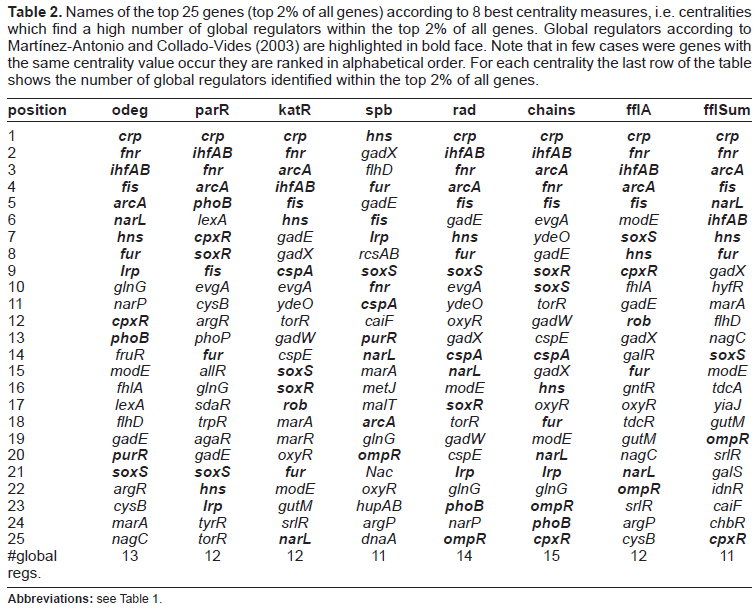
The top 25 genes (top 2% of all genes) according to the eight best centrality measures (i.e. the centrality measures which identify the highest number of global regulators within the top 2% of all genes) are shown in Table 2. In total 18 global regulators have been identified by Martínez-Antonio and Collado-Vides (2003). All different centrality measures shown in Table 2 are able to identify more than 50% of the global regulators within the top 2% of the ranked genes. For example, shortest path betweenness finds 11 global regulators and motif-based centrality with the chain motif class is able to identify 15 global regulators.
Names of the top 25 genes (top 2% of all genes) according to 8 best centrality measures, i.e. centralities which find a high number of global regulators within the top 2% of all genes. Global regulators according to Martínez-Antonio and Collado-Vides (2003) are highlighted in bold face. Note that in few cases were genes with the same centrality value occur they are ranked in alphabetical order. For each centrality the last row of the table shows the number of global regulators identified within the top 2% of all genes.
It should be also noted that for nearly all centrality measures the top 5 positions are occupied by global regulators. However, all centralities result in different rankings even for global regulators which are often ranked very high. For example, the gene
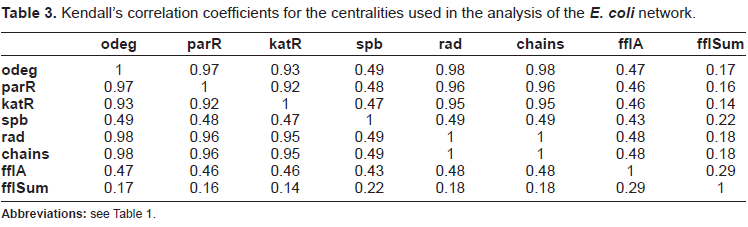
Correlation coefficients are a valid measure to show that centralities do not coincide. Table 3 shows the pairwise Kendall's correlation coefficients for the centralities used in Table 2. From these centralities only a few correlate with a coefficient above 0.9 to other centralities. These are out-degree, PageRank, Katz status index, radiality and the motif-based centrality with chain classes (chain). The centralities based on the FFL motif and shortest-path betweenness do correlate only with correlation coefficients less than 0.9 to other centralities.
Kendall's correlation coefficients for the centralities used in the analysis of the
For the five centralities with a correlation coefficient above 0.9 these high coefficients can easily be explained: 1101 out of 1250 (88.08%) vertices have an out-degree of zero. All these vertices are assigned the same centrality value of nearly zero for the Katz status index and the PageRank centrality, and the value zero for the radiality and the motif-based centrality with chain classes. Therefore, the comparison of correlations between all centrality values is not feasible for the complete vector of centralities: all five centralities rank these 1101 vertices into the same group.
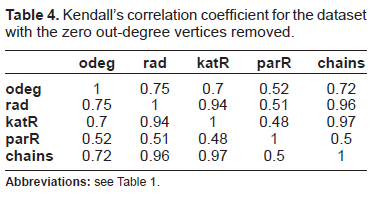
Table 4 shows the pairwise correlation coefficients for the centrality values of the vertices which have a non-zero out-degree. These coefficients show a different picture: all five centralities do rank the remaining 149 genes differently, only the centrality radiality and Katz status index archive a considerable high correlation to each other and to the motif-based centrality with chain classes.
Kendall's correlation coefficient for the dataset with the zero out-degree vertices removed.
In conclusion, the centralities applied to the GRN rank the genes differently and the motif-based centrality with chain classes is able to rank the highest number of interesting genes (global regulators) within the top 2% of all genes. The chain centrality identifies 15 out of 18 global regulators (83%) identified by Martínez-Antonio and Collado-Vides (2003) and outperforms the other centralities used.
Discussion
To investigate large biological networks different analysis methods have been developed, and centrality analysis is a particularly useful method to analyze the structure of these networks. In this paper we discussed and compared different centrality measures and applied them to a gene regulatory network of
The comparison of the pairwise correlation coefficients and the analysis of the rankings of the top 25 genes show that the motif-based centralities, in particular with the chain motif class, produce rankings different to the rankings computed by existing centralities, and that these rankings show interesting features of the gene regulatory network under analysis.
